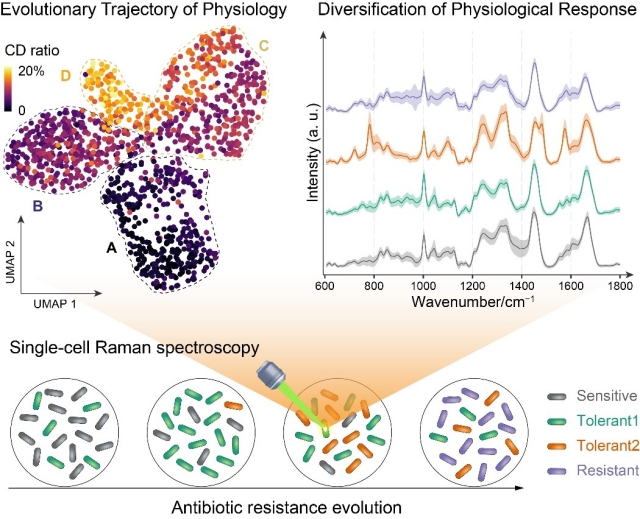
Understanding evolution of antibiotic resistance is vital for containing its global spread. Yet our ability to in situ track highly heterogenous and dynamic evolution is very limited. Here, we present a new single-cell approach integrating D2O-labeled Raman spectroscopy, advanced multivariate analysis, and genotypic profiling to in situ track physiological evolution trajectory toward resistance. Physiological diversification of individual cells from isogenic population with cyclic ampicillin treatment is captured. Advanced multivariate analysis of spectral changes classifies all individual cells into four subsets of sensitive, intrinsic tolerant, evolved tolerant and resistant. Remarkably, their dynamic shifts with evolution are depicted and spectral markers of each state are identified. Genotypic analysis validates the phenotypic shift and provides insights into the underlying genetic basis. The new platform advances rapid phenotyping resistance evolution and guides evolution control.